Volumetric Spleen Segmentation via Attention U-Net
Overview
Engineered an end-to-end 3D imaging pipeline to segment the spleen from highly variable CT scans. To scientifically validate the architecture, I developed and benchmarked five distinct models against a baseline standard U-Net.
Clinical Significance
In clinical practice, a model that simply outputs a prediction is insufficient; clinicians need to know *when* the model is unsure. By building a dual-head Bayesian Neural Network into this pipeline, the model generates diagnostic heatmaps that isolate Epistemic uncertainty (model ignorance) from Aleatoric uncertainty (CT acquisition artifacts).
Model Benchmarking
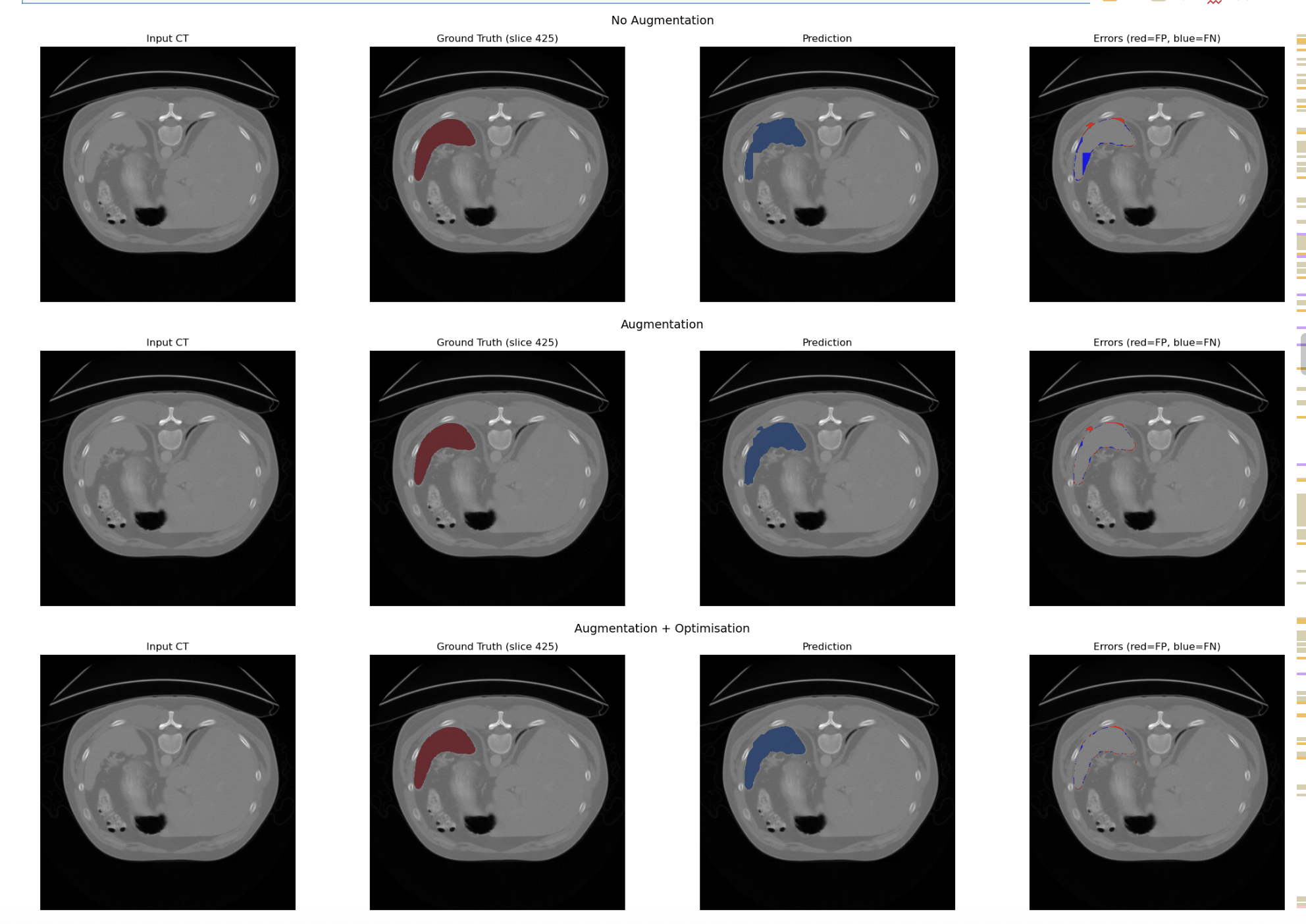
- Baseline U-Net: Standard 3D U-Net without attention gating.
- Attention U-Net: Integrated Attention Gates to suppress background anatomy.
- Attention U-Net with Data Augmentation: Random Affine and Elastic Deformation of data introduced to mimic normal anatomical variation/scanner artifacts
- Attention U-Net with Data Augmentation and Optimisation: Implemented AdamW optimiser and Cosine Annealing Scheduler
- Attention U-Net with Augmentation Consistency:Augmentation Consistency Technique (where volumes are variabily augmentated and passed through the same model for prediction) used to futher enhance model's performance
- Bayesian Attention U-Net: Upgraded to quantify uncertainty and output Aleatoric and Epistemic heatmaps.
Pre-processing
The dataset that I was working with for this project was relatively small (41 CT volumes) thus I decided to split my training into 35 volumes for training, 3 for validation and 3 for testing.
As there were inconsistencies in resolutions between different views (axial, coronal and saggital), I used SimpleITK to ensure isometricity of all the views first to prevent the model from learning from distorted anatomy
Given the small dataset, I was able to preload all of the volumes into memory which significantly reduced data-loading bottlenecks and accelerate computation.
The isometric volumes and their corresponding ground-truth labels were divided into patches. Preventing the network from being overwhelmed by emptybackground voxels, I used a balanced patch sampling strategy. Patches containing spleen tissue were randomly shuffled and paired with background only patches in a 1:1 ratio, ensuring the model received sufficient foreground signal during training.
Network Architecture
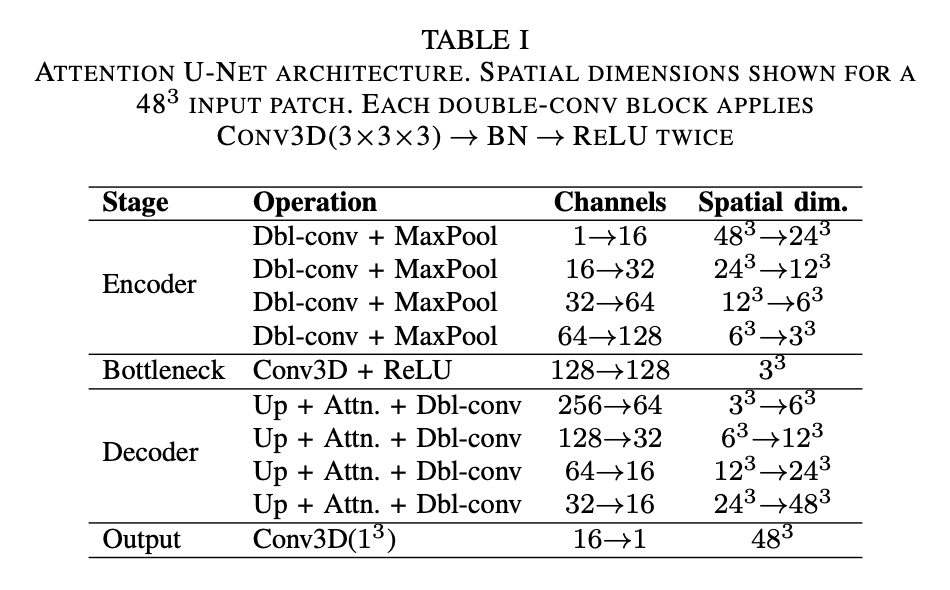
The base network chosen for this segmentation task is an Attention U-Net, modelled after the architecture proposed by Oktay et al. (2018). This network processes 3D image patches and can be conceptually divided into three components:
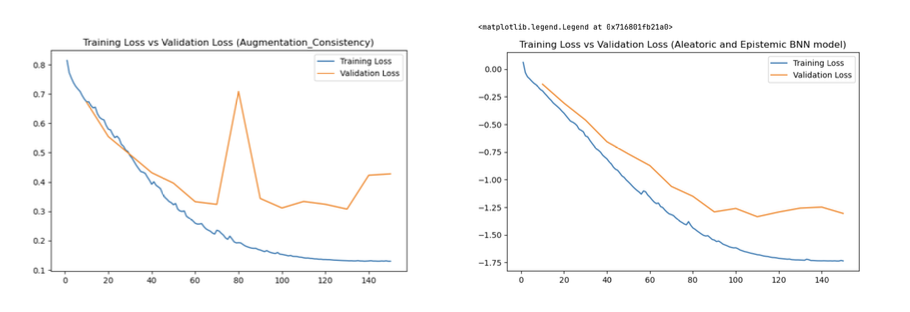
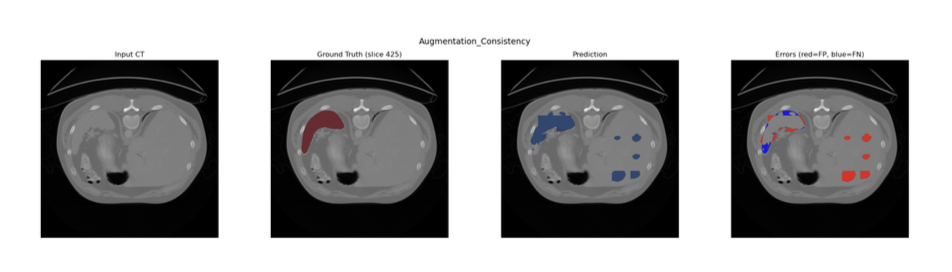
Augmentation Consistency Model (Model D)
To leverage the abundance of unlabelled image volumes, an augmentation consistency framework was integrated. The training loop was partitioned into two streams:
1. Supervised - Labelled patches were passed through the network without additional augmentation
2. Unsupervised - Two independent augmentations were applied to the same unlabelled patch. Both augmented views were passed through the current model under gradient isolation; the MSE between the two resulting predictions was computed as the consistency loss and scaled by λ = 0.01 before backpropagation.
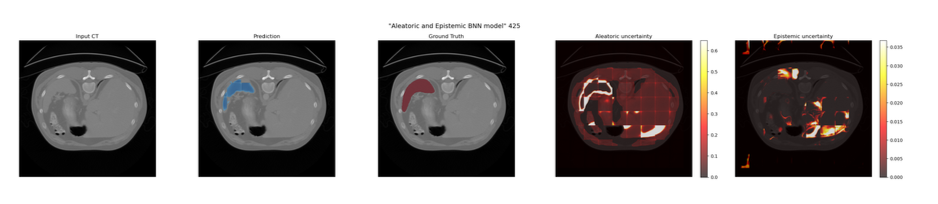
Bayesian Neural Network Model (Model E)
Epistemic Uncertainty: A 3D Dropout layer (p=0.5) was introduced at the bottleneck. During inference, Monte Carlo (MC) Dropout was utilised by keeping the model in training mode and executing 20 forward passes. Epistemic uncertainty was calculated as the variance of predictions across these passes.
Aleatoric Uncertainty: The network was split into a dual-head decoder after the bottleneck. The Prediction Head yielded the class probability map, while the Variance Head output a log-variance map (predicting the logarithm ensures positive variance).
Results
Model A,B and C
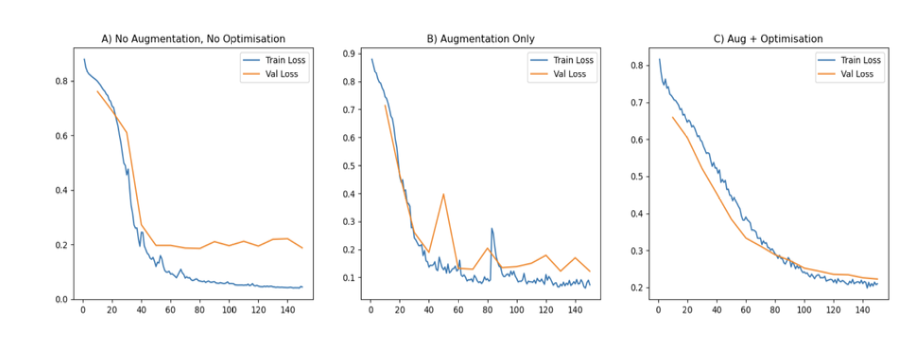
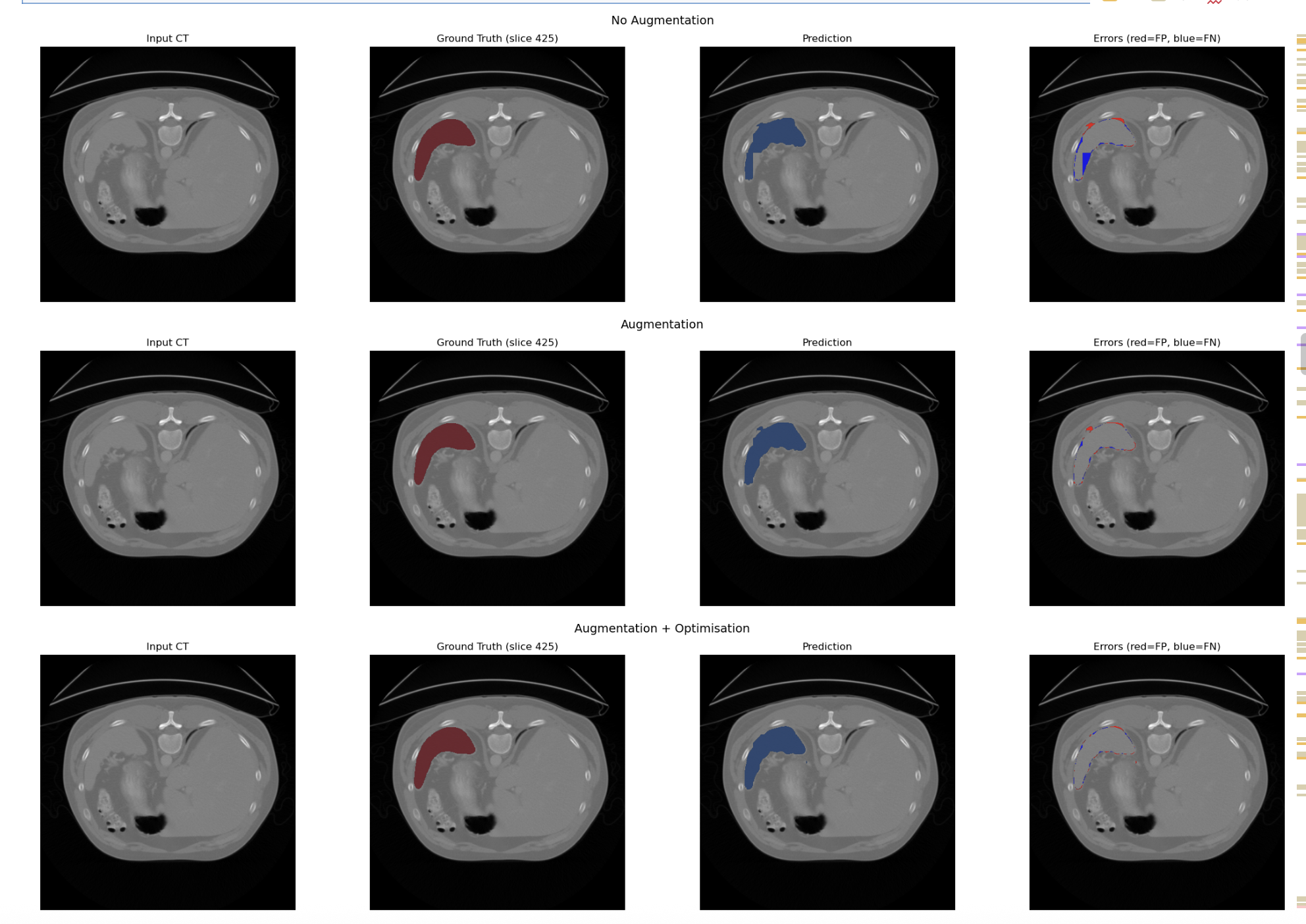
Model D and E